“Most of what we do that resonates with the public is to get weird, unexpected microbes out of the environment.” The speaker is Michelle O’Malley, assistant professor in the Department of Chemical Engineering at the UCSB College of Engineering, and the environment she’s talking about is, often, the gut of a large herbivore, such as a cow, a horse, or a zoo animal. That’s where she and her research colleagues have discovered organisms that can be engineered to serve human needs.
In a paper titled “A Parts List for Fungal Cellulosomes Revealed by Comparative Genomics” and published today in Nature Microbiology, O’Malley and more than twenty co-investigators describe a new complex of enzymes discovered in herbivore gut fungi that may have applications in sustainable fuels and chemicals.
Enzymes are the powerhouses behind biological chemistry, and the fungi discovered by O’Malley’s group — like Anaeromyces robustus (named after the gray whale, partially based on how it looks under the microscope) — has unusual and desirable characteristics, particularly the ability to transform lignocellulose from plants into sugars.
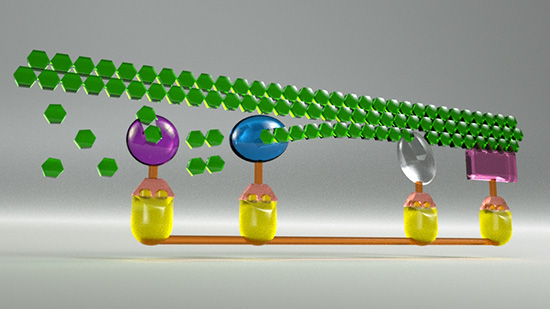
The illustration above depicts a chain of repeating green glucose molecules in cellulose being broken down by A. robustus. The rodlike horizontal relement at the bottom and the yellow parts rising vertically from it represent the scaffoldin protein. Topping them, the bronze-colored trapezoids with vertical extenders are the dockerin protein domains, which allow the various types of enzymes (shown as gemlike elements) to work together in assembly-line fashion to break down the cellulose.
In the unique structure of A. robustus, individual enzymes are arrayed as a kind of large protein mass or scaffold, called a cellulosome, so that they stick to each other and accumulate, somewhat as Legos do. A structure on the enzymes — a dockerin — allows the enzyme to plug into the scaffolding mechanism. The result, O’Malley explained, is that the entire structure “kind of glues the enzymes together for maximum impact to break down the non-food parts of plants.” This process stands in stark contrast to how industry accomplishes the same thing, by relying on free-floating enzyme mixtures to break down biomass.
That mechanism could make a difference in cellulosic biofuel plants, which produce ethanol from waste products — corn husks and cobs — rather than edible kernels, a major advance in addressing the tradeoff of using agricultural land to grow corn for fuel rather than for food. “From our preliminary evidence, the enzymes we’re studying have the potential to be better for industrial use than the ones being used now,” said Sean Gilmore, a PhD student in O’Malley’s lab and a co-author on the paper, who also identified the new enzymes while examining the genomes of the new fungi.
In current cellulosic biofuel production, O’Malley explained, “You need to have several different ‘flavors’ of enzymes to get the end product you want. Each enzyme has a different job. Those enzymes are added as a kind of cocktail mixture, and they run into each other randomly to create the necessary reactants. What’s different about the structures we’ve identified is that the enzymes are tethered, so the reactions and their products are moved along an assembly line, expanding their activity for maximum impact rather than happening by luck.”
Similarly functioning structures are found in bacteria, but they are quite different in that environment. “The principal is the same, but the machinery is completely different,” O’Malley said.
“It suggests that they evolved ndependent of each other, toward the same purpose of breaking down cellulosic material,” Gilmore added. “They started from different places evolutionarily but occupy the same kind of environment, so they developed similar structures in different ways. Nature has kind of pushed them that way for a reason; they’re filling a specific niche.”
This is the first report to describe cellulosome structures in fungi, and to note that the fungi have “stolen” some of the parts from bacteria, through a process called “horizontal gene transfer.” O’Malley described it as similar to “living with a roommate and picking up some of their habits. The fungi and the bacteria evolved these machines differently, but along the way the fungi took something they liked from the bacteria. That, too, is unusual, especially given that on the tree of life, even primitive anaerobic fungi, like the three that underwent genome sequencing in this study, are evolutionarily much closer to humans than to bacteria. That makes this instance of horizontal gene transfer “novel, because it’s a long jump,” O’Malley said.
Understanding the newly identified complexes gives us a better understanding of how cellulose is converted to sugar. “From a biotech perspective, that gives us a recipe to try to build similar complexes that could then be engineered to more efficiently break down widely available crop waste,” O’Malley noted. “It’s the whole idea of value-added production: you take something that people would just burn before and then use these enzyme machines to extract sugar from it, which is fed to microbes, which can then make things for us. Further, knowledge of these fungal complexes could be applied to engineer completely synthetic assembly lines to build, for instance, novel chemicals and pharmaceuticals.”
One of the key processes in the research was comparing the genome assemblies of the fungal enzymes to those of other biomass-degrading microbes, made possible only by recent breakthroughs in genome sequencing techniques,
Gilmore recalls having the ‘Wow’ moment while looking at the sequences rather than the physical protein: “When we got the sequence back, the first thing I did was to sort by size, and at the top you could see a large, uncharacterized protein. I thought, That can’t be right.”
“We had thought to look for something like this, because we knew the enzymes had to assemble onto something,” O’Malley added, “but we didn’t know what to look for other than that the thing should probably be big and repetitious in structure.
“It’s cool to find things, and it’s rare for engineers,” O’Malley added. I’m very proud of the fact that our research tries to balance efforts in discovery and engineering. And working on unusual microbes offers a route to do that.”

Assistant Professor Michelle O'Malley
